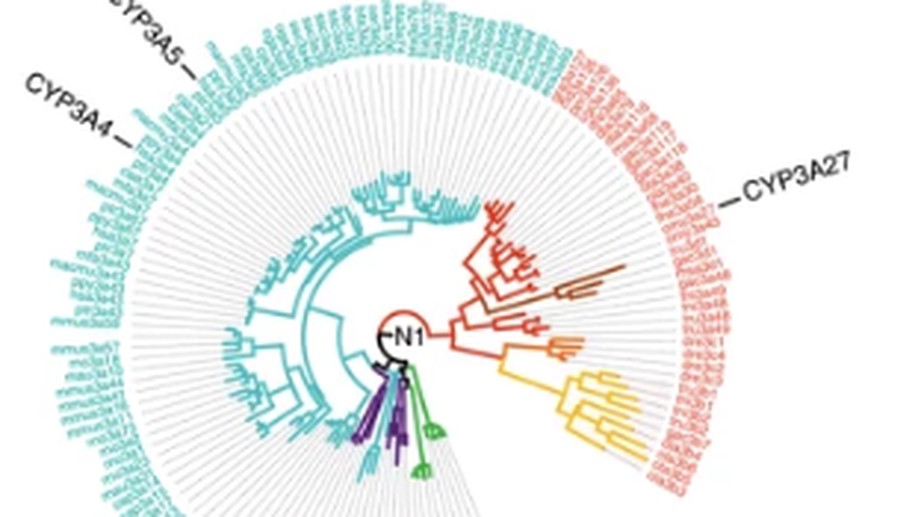
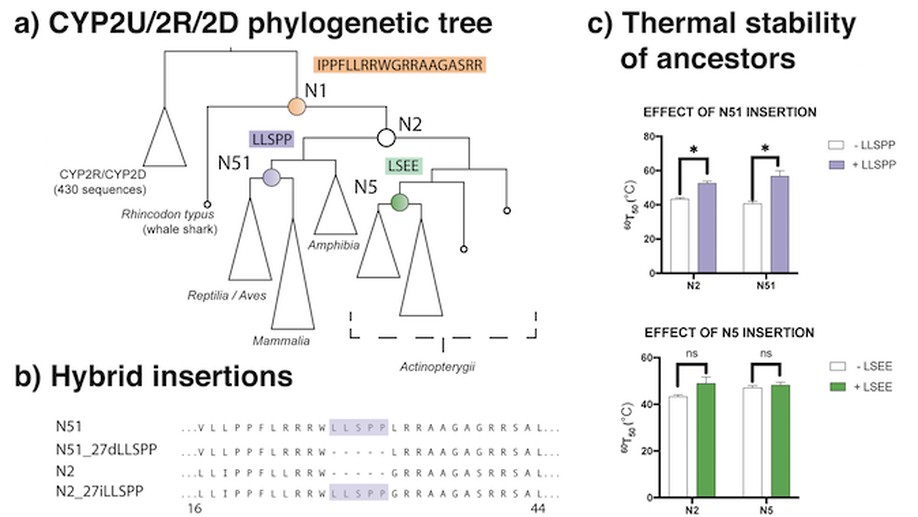
GRASP-suite is a collection of tools and tutorials to perform and analyse ancestral sequence reconstruction.
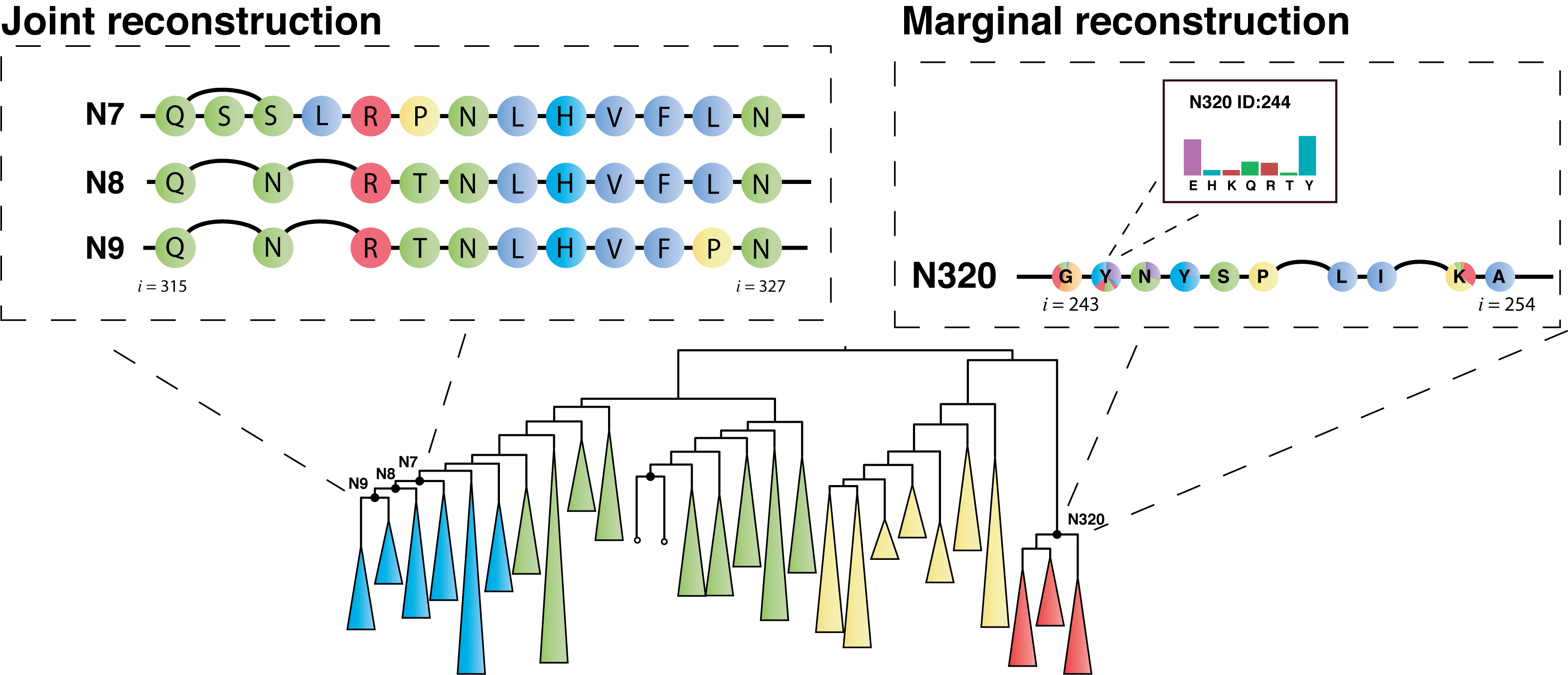
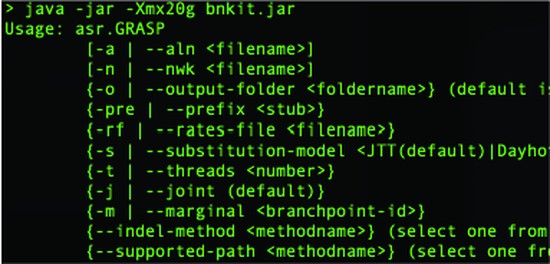
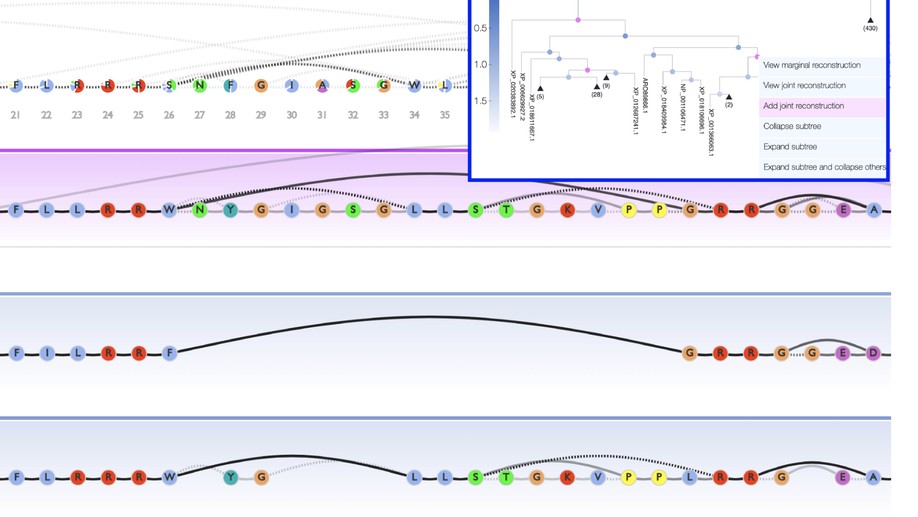
Central to the GRASP-suite is Graphical Representation of Ancestral Sequence Predictions (GRASP), an ancestral sequence reconstruction tool capable of performing maximum likelihood analysis on very large data sets.